Transverse Relaxation-Optimized SpectroscopY (TROSY) used in combination with various isotope labeling techniques enables NMR studies of large proteins and biomolecular complexes. Stereo Array Isotope Labeling (SAIL) is a strategy developed by Prof. Kainosho in 2006 that exploits stereo-selective 13C and 2H labeling of amino acids to limit spin diffusion, reduce signal overlapping and allow stereospecific assignment of prochiral groups.
The SAIL-TROSY approach combines the SAIL strategy with TROSY NMR techniques for structural investigations on biomolecular systems of molecular weights up to 1,000 kDa. Originally limited to the observation of backbone amides (15NH-TROSY) and side-chain methyls (methyl 13C-TROSY), the SAIL-TROSY approach has been extended to the generation of narrow NMR linewidth for aromatics (aromatic-13CH-TROSY) or aliphatic methylene and methyl groups (aliphatic 13CH-TROSY).
As an example, the SAIL-TROSY approach was recently applied for the observation of aromatic ring 13CH signals of the 82 kDa single domain protein Malate Synthase G (MSG). The protein was deuterated, uniformly labeled with 15N and selectively labeled with [δ1,ε3,η2]-SAIL Trp, allowing the obtention of 36 well-resolved cross peaks for the δ1, ε3, an η2 CHs for the 12 Trp residues of MSG and complete signal assignment (Figure below).
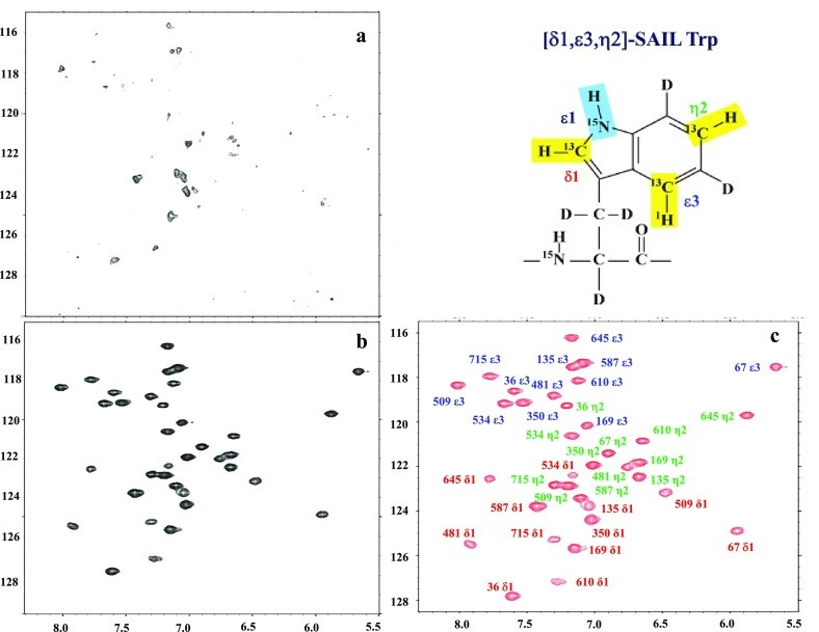
Aromatic regions of 800 MHz 2D 1H–13C correlation NMR spectra of 80 kDa Malate Synthase G (MSG) protein selectively labeled with [δ1,ε3,η2]-SAIL Trp, and otherwise uniformly labeled with deuterium and 15N. a F1-decoupled HSQC; b Aromatic 13CH TROSY; c Aromatic 13CH TROSY with signal assignment. Adapted from Kainosho et al., J. Biomol. NMR, 2018.
All SAIL approaches can be employed for structural and dynamics investigations on proteins produced in bacteria or insect cell and is particularly adapted to cell-free expression systems.

Cortec Isotope is pleased to offer a wide selection of SAIL amino acids for Methyl 13C TROSY and Aromatic 13C TROSY applications available for direct use. Custom-made SAIL amino acids mixtures containing selectively methyl, aromatic and deuterated labeled amino acids are available on request.
Cortec Isotope proposes to help you label your protein. For inquiry, please contact our labeled protein expression platform.